| Multiple Linear Regression - Estimated Regression Equation |
| A[t] = -59199.6 + 8.25972B[t] + 181.499C[t] -92.2104D[t] + 785.242E[t] + 42.8427F[t] + 5.04409G[t] + 50.1865H[t] -1829.25I[t] -1846.08J[t] + 107.804K[t] + 26.5595L[t] + 0.742319M[t] + 10.2978N[t] -6.70532O[t] + e[t] |
| Multiple Linear Regression - Ordinary Least Squares | |||||
| Variable | Parameter | S.D. | T-STAT H0: parameter = 0 | 2-tail p-value | 1-tail p-value |
| (Intercept) | -5.92e+04 | 1.585e+04 | -3.7340e+00 | 0.0002705 | 0.0001353 |
| B | +8.26 | 6.63 | +1.2460e+00 | 0.2149 | 0.1074 |
| C | +181.5 | 91.25 | +1.9890e+00 | 0.04859 | 0.0243 |
| D | -92.21 | 47.2 | -1.9530e+00 | 0.0527 | 0.02635 |
| E | +785.2 | 232.6 | +3.3760e+00 | 0.0009471 | 0.0004735 |
| F | +42.84 | 136.9 | +3.1290e-01 | 0.7548 | 0.3774 |
| G | +5.044 | 1.605 | +3.1420e+00 | 0.002035 | 0.001017 |
| H | +50.19 | 18.73 | +2.6790e+00 | 0.008242 | 0.004121 |
| I | -1829 | 1083 | -1.6890e+00 | 0.09331 | 0.04665 |
| J | -1846 | 781.9 | -2.3610e+00 | 0.01956 | 0.009781 |
| K | +107.8 | 77.3 | +1.3950e+00 | 0.1653 | 0.08265 |
| L | +26.56 | 16.53 | +1.6060e+00 | 0.1104 | 0.05519 |
| M | +0.7423 | 0.568 | +1.3070e+00 | 0.1933 | 0.09666 |
| N | +10.3 | 155.8 | +6.6090e-02 | 0.9474 | 0.4737 |
| O | -6.705 | 140 | -4.7890e-02 | 0.9619 | 0.4809 |
| Multiple Linear Regression - Regression Statistics | |
| Multiple R | 0.9223 |
| R-squared | 0.8507 |
| Adjusted R-squared | 0.8361 |
| F-TEST (value) | 58.59 |
| F-TEST (DF numerator) | 14 |
| F-TEST (DF denominator) | 144 |
| p-value | 0 |
| Multiple Linear Regression - Residual Statistics | |
| Residual Standard Deviation | 2379 |
| Sum Squared Residuals | 8.152e+08 |
| Menu of Residual Diagnostics | |
| Description | Link |
| Histogram | Compute |
| Central Tendency | Compute |
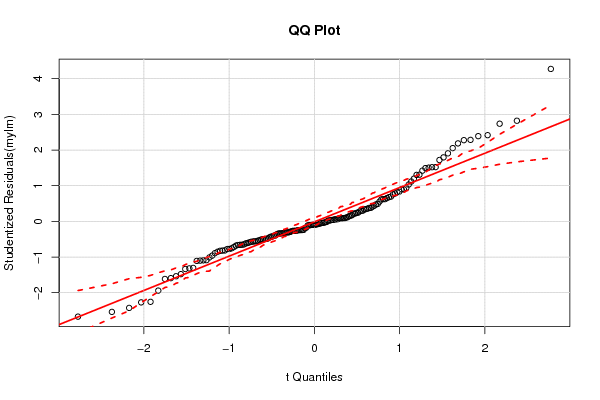
| QQ Plot | Compute |
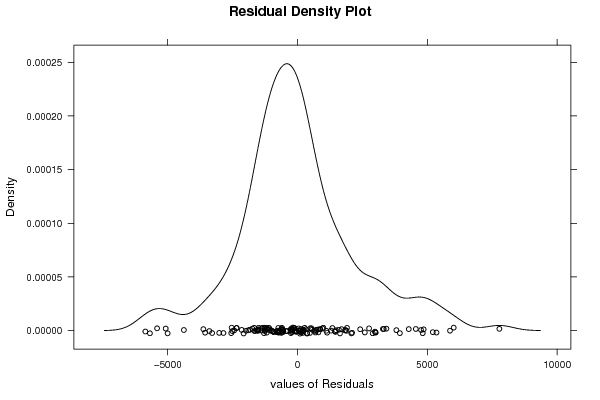
| Kernel Density Plot | Compute |
| Skewness/Kurtosis Test | Compute |
| Skewness-Kurtosis Plot | Compute |
| Harrell-Davis Plot | Compute |
| Bootstrap Plot -- Central Tendency | Compute |
| Blocked Bootstrap Plot -- Central Tendency | Compute |
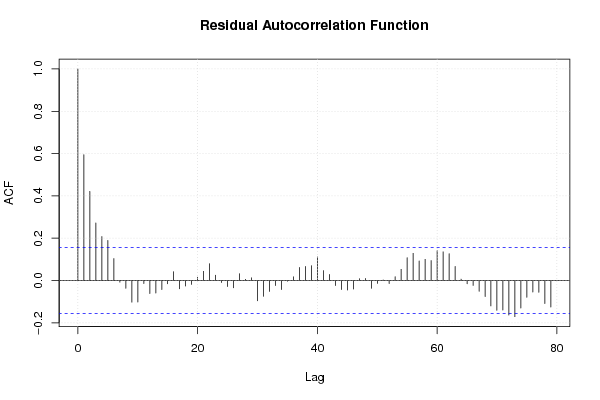
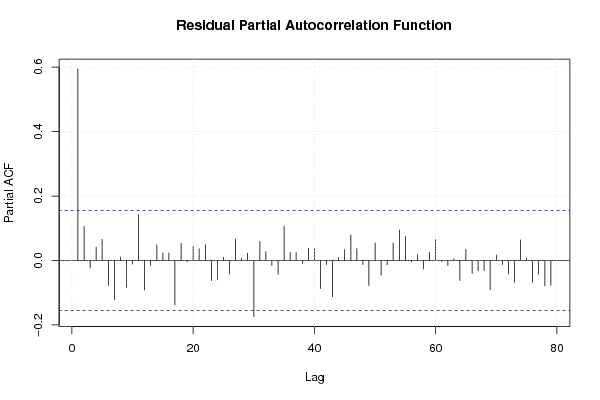
| (Partial) Autocorrelation Plot | Compute |
| Spectral Analysis | Compute |
| Tukey lambda PPCC Plot | Compute |
| Box-Cox Normality Plot | Compute |
| Summary Statistics | Compute |
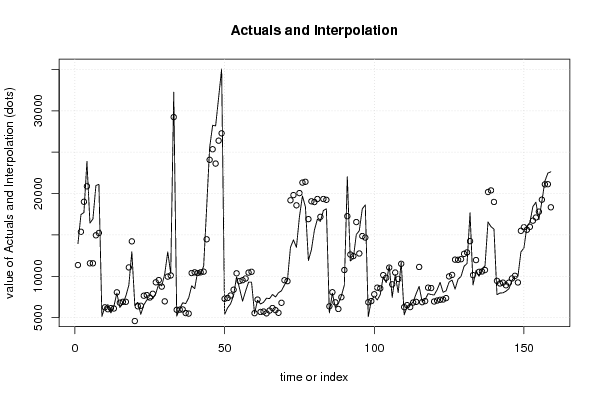
| Multiple Linear Regression - Actuals, Interpolation, and Residuals | |||
| Time or Index | Actuals | Interpolation Forecast | Residuals Prediction Error |
| 1 | 1.395e+04 | 1.136e+04 | 2594 |
| 2 | 1.745e+04 | 1.537e+04 | 2077 |
| 3 | 1.771e+04 | 1.9e+04 | -1289 |
| 4 | 2.388e+04 | 2.088e+04 | 3000 |
| 5 | 1.643e+04 | 1.157e+04 | 4860 |
| 6 | 1.692e+04 | 1.157e+04 | 5355 |
| 7 | 2.097e+04 | 1.495e+04 | 6018 |
| 8 | 2.11e+04 | 1.523e+04 | 5876 |
| 9 | 5151 | 326.5 | 4825 |
| 10 | 6295 | 6246 | 48.81 |
| 11 | 6575 | 6015 | 560.1 |
| 12 | 5572 | 6145 | -572.8 |
| 13 | 6377 | 6102 | 275 |
| 14 | 7957 | 8060 | -103 |
| 15 | 6229 | 6800 | -571.3 |
| 16 | 6692 | 6911 | -219.2 |
| 17 | 7609 | 6911 | 697.8 |
| 18 | 8921 | 1.107e+04 | -2149 |
| 19 | 1.296e+04 | 1.422e+04 | -1256 |
| 20 | 6479 | 4590 | 1889 |
| 21 | 6855 | 6372 | 482.7 |
| 22 | 5399 | 6434 | -1035 |
| 23 | 6529 | 7648 | -1119 |
| 24 | 7129 | 7729 | -599.9 |
| 25 | 7295 | 7430 | -134.6 |
| 26 | 7295 | 7883 | -587.8 |
| 27 | 7895 | 9271 | -1376 |
| 28 | 9095 | 9539 | -443.5 |
| 29 | 8845 | 8747 | 98.44 |
| 30 | 1.03e+04 | 6969 | 3326 |
| 31 | 1.294e+04 | 9960 | 2985 |
| 32 | 1.034e+04 | 1.01e+04 | 243.2 |
| 33 | 3.225e+04 | 2.924e+04 | 3013 |
| 34 | 5195 | 5948 | -753.1 |
| 35 | 6095 | 5962 | 133.1 |
| 36 | 6795 | 5987 | 807.9 |
| 37 | 6695 | 5553 | 1142 |
| 38 | 7395 | 5487 | 1908 |
| 39 | 8845 | 1.039e+04 | -1545 |
| 40 | 8495 | 1.048e+04 | -1983 |
| 41 | 1.06e+04 | 1.039e+04 | 204.8 |
| 42 | 1.024e+04 | 1.048e+04 | -232.8 |
| 43 | 1.124e+04 | 1.055e+04 | 691.6 |
| 44 | 1.828e+04 | 1.447e+04 | 3809 |
| 45 | 2.555e+04 | 2.407e+04 | 1480 |
| 46 | 2.825e+04 | 2.535e+04 | 2896 |
| 47 | 2.818e+04 | 2.362e+04 | 4559 |
| 48 | 3.16e+04 | 2.639e+04 | 5214 |
| 49 | 3.506e+04 | 2.728e+04 | 7779 |
| 50 | 5389 | 7282 | -1893 |
| 51 | 6189 | 7372 | -1183 |
| 52 | 6669 | 7674 | -1005 |
| 53 | 7689 | 8374 | -684.9 |
| 54 | 9959 | 1.036e+04 | -399.1 |
| 55 | 8499 | 9419 | -920.4 |
| 56 | 6989 | 9524 | -2535 |
| 57 | 8189 | 9726 | -1537 |
| 58 | 9279 | 1.044e+04 | -1164 |
| 59 | 9279 | 1.054e+04 | -1263 |
| 60 | 5499 | 5538 | -39.08 |
| 61 | 7099 | 7181 | -82.02 |
| 62 | 6649 | 5684 | 964.6 |
| 63 | 6849 | 5736 | 1113 |
| 64 | 7349 | 5518 | 1831 |
| 65 | 7299 | 5851 | 1448 |
| 66 | 7799 | 6160 | 1639 |
| 67 | 7499 | 5902 | 1597 |
| 68 | 7999 | 5583 | 2416 |
| 69 | 8249 | 6794 | 1455 |
| 70 | 8949 | 9520 | -570.9 |
| 71 | 9549 | 9409 | 140.1 |
| 72 | 1.35e+04 | 1.918e+04 | -5685 |
| 73 | 1.44e+04 | 1.981e+04 | -5409 |
| 74 | 1.35e+04 | 1.858e+04 | -5076 |
| 75 | 1.72e+04 | 2.005e+04 | -2853 |
| 76 | 1.97e+04 | 2.132e+04 | -1621 |
| 77 | 1.84e+04 | 2.141e+04 | -3016 |
| 78 | 1.19e+04 | 1.69e+04 | -5004 |
| 79 | 1.32e+04 | 1.906e+04 | -5861 |
| 80 | 1.558e+04 | 1.897e+04 | -3394 |
| 81 | 1.69e+04 | 1.934e+04 | -2438 |
| 82 | 1.663e+04 | 1.718e+04 | -550.9 |
| 83 | 1.795e+04 | 1.934e+04 | -1388 |
| 84 | 1.815e+04 | 1.924e+04 | -1089 |
| 85 | 5572 | 6364 | -791.8 |
| 86 | 7957 | 8068 | -111.3 |
| 87 | 6229 | 6850 | -620.8 |
| 88 | 6692 | 6047 | 644.7 |
| 89 | 7609 | 7468 | 141.3 |
| 90 | 8921 | 1.075e+04 | -1833 |
| 91 | 2.202e+04 | 1.726e+04 | 4760 |
| 92 | 1.185e+04 | 1.263e+04 | -779.1 |
| 93 | 1.217e+04 | 1.243e+04 | -264.8 |
| 94 | 1.504e+04 | 1.655e+04 | -1511 |
| 95 | 1.551e+04 | 1.275e+04 | 2757 |
| 96 | 1.815e+04 | 1.486e+04 | 3291 |
| 97 | 1.862e+04 | 1.468e+04 | 3945 |
| 98 | 5118 | 6857 | -1739 |
| 99 | 7053 | 6995 | 57.67 |
| 100 | 7603 | 7826 | -223.1 |
| 101 | 7126 | 8619 | -1493 |
| 102 | 7775 | 8535 | -759.9 |
| 103 | 9960 | 1.014e+04 | -176.3 |
| 104 | 9233 | 9815 | -581.8 |
| 105 | 1.126e+04 | 1.105e+04 | 210.4 |
| 106 | 7463 | 9028 | -1565 |
| 107 | 1.02e+04 | 1.045e+04 | -254.2 |
| 108 | 8013 | 9674 | -1661 |
| 109 | 1.169e+04 | 1.15e+04 | 189.7 |
| 110 | 5348 | 6267 | -918.8 |
| 111 | 6338 | 6510 | -171.7 |
| 112 | 6488 | 6276 | 211.7 |
| 113 | 6918 | 6827 | 90.81 |
| 114 | 7898 | 6903 | 995 |
| 115 | 8778 | 1.112e+04 | -2344 |
| 116 | 6938 | 6867 | 71.38 |
| 117 | 7198 | 6999 | 198.7 |
| 118 | 7898 | 8619 | -721 |
| 119 | 7788 | 8578 | -789.9 |
| 120 | 7738 | 6948 | 790.5 |
| 121 | 8358 | 7064 | 1294 |
| 122 | 9258 | 7155 | 2103 |
| 123 | 8058 | 7186 | 872 |
| 124 | 8238 | 7363 | 875.5 |
| 125 | 9298 | 9984 | -685.9 |
| 126 | 9538 | 1.016e+04 | -622.4 |
| 127 | 8449 | 1.201e+04 | -3559 |
| 128 | 9639 | 1.199e+04 | -2348 |
| 129 | 9989 | 1.206e+04 | -2074 |
| 130 | 1.12e+04 | 1.271e+04 | -1510 |
| 131 | 1.155e+04 | 1.289e+04 | -1336 |
| 132 | 1.767e+04 | 1.424e+04 | 3424 |
| 133 | 8948 | 1.014e+04 | -1188 |
| 134 | 1.07e+04 | 1.196e+04 | -1260 |
| 135 | 9988 | 1.053e+04 | -542.3 |
| 136 | 1.09e+04 | 1.057e+04 | 324.9 |
| 137 | 1.125e+04 | 1.075e+04 | 495.8 |
| 138 | 1.656e+04 | 2.018e+04 | -3625 |
| 139 | 1.6e+04 | 2.037e+04 | -4376 |
| 140 | 1.569e+04 | 1.898e+04 | -3289 |
| 141 | 7775 | 9441 | -1666 |
| 142 | 7975 | 9131 | -1156 |
| 143 | 7995 | 9225 | -1230 |
| 144 | 8195 | 8915 | -719.7 |
| 145 | 8495 | 9233 | -737.5 |
| 146 | 9495 | 9732 | -236.8 |
| 147 | 9995 | 1.005e+04 | -58.5 |
| 148 | 9980 | 9248 | 731.7 |
| 149 | 1.294e+04 | 1.549e+04 | -2553 |
| 150 | 1.342e+04 | 1.592e+04 | -2509 |
| 151 | 1.598e+04 | 1.562e+04 | 366 |
| 152 | 1.652e+04 | 1.597e+04 | 540.1 |
| 153 | 1.842e+04 | 1.672e+04 | 1701 |
| 154 | 1.895e+04 | 1.71e+04 | 1850 |
| 155 | 1.684e+04 | 1.78e+04 | -959.5 |
| 156 | 1.904e+04 | 1.926e+04 | -210.5 |
| 157 | 2.148e+04 | 2.111e+04 | 377.1 |
| 158 | 2.247e+04 | 2.112e+04 | 1346 |
| 159 | 2.262e+04 | 1.834e+04 | 4287 |
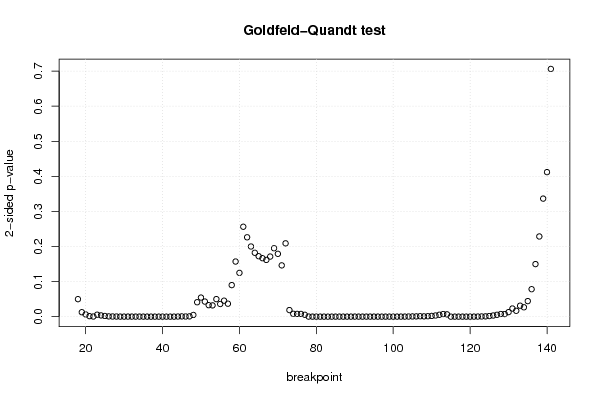
| Goldfeld-Quandt test for Heteroskedasticity | |||
| p-values | Alternative Hypothesis | ||
| breakpoint index | greater | 2-sided | less |
| 18 | 0.02497 | 0.04994 | 0.975 |
| 19 | 0.006353 | 0.01271 | 0.9936 |
| 20 | 0.003197 | 0.006393 | 0.9968 |
| 21 | 0.0007184 | 0.001437 | 0.9993 |
| 22 | 0.0002077 | 0.0004154 | 0.9998 |
| 23 | 0.002773 | 0.005547 | 0.9972 |
| 24 | 0.001789 | 0.003578 | 0.9982 |
| 25 | 0.0009568 | 0.001914 | 0.999 |
| 26 | 0.0003917 | 0.0007835 | 0.9996 |
| 27 | 0.0002211 | 0.0004422 | 0.9998 |
| 28 | 0.000169 | 0.0003379 | 0.9998 |
| 29 | 0.0001102 | 0.0002204 | 0.9999 |
| 30 | 4.676e-05 | 9.351e-05 | 1 |
| 31 | 9.648e-05 | 0.000193 | 0.9999 |
| 32 | 8.893e-05 | 0.0001779 | 0.9999 |
| 33 | 6.419e-05 | 0.0001284 | 0.9999 |
| 34 | 9.6e-05 | 0.000192 | 0.9999 |
| 35 | 5.171e-05 | 0.0001034 | 0.9999 |
| 36 | 3.704e-05 | 7.409e-05 | 1 |
| 37 | 1.708e-05 | 3.415e-05 | 1 |
| 38 | 8.78e-06 | 1.756e-05 | 1 |
| 39 | 4.707e-06 | 9.413e-06 | 1 |
| 40 | 2.272e-06 | 4.544e-06 | 1 |
| 41 | 7.54e-06 | 1.508e-05 | 1 |
| 42 | 4.937e-06 | 9.873e-06 | 1 |
| 43 | 6.091e-06 | 1.218e-05 | 1 |
| 44 | 0.0001879 | 0.0003757 | 0.9998 |
| 45 | 0.0003566 | 0.0007133 | 0.9996 |
| 46 | 0.000224 | 0.000448 | 0.9998 |
| 47 | 0.0004366 | 0.0008733 | 0.9996 |
| 48 | 0.002402 | 0.004804 | 0.9976 |
| 49 | 0.02055 | 0.0411 | 0.9795 |
| 50 | 0.02721 | 0.05442 | 0.9728 |
| 51 | 0.02153 | 0.04305 | 0.9785 |
| 52 | 0.01637 | 0.03275 | 0.9836 |
| 53 | 0.01609 | 0.03218 | 0.9839 |
| 54 | 0.02501 | 0.05003 | 0.975 |
| 55 | 0.01804 | 0.03608 | 0.982 |
| 56 | 0.02275 | 0.04549 | 0.9773 |
| 57 | 0.01847 | 0.03694 | 0.9815 |
| 58 | 0.04496 | 0.08992 | 0.955 |
| 59 | 0.07858 | 0.1572 | 0.9214 |
| 60 | 0.06245 | 0.1249 | 0.9375 |
| 61 | 0.1282 | 0.2564 | 0.8718 |
| 62 | 0.1132 | 0.2263 | 0.8868 |
| 63 | 0.1 | 0.2001 | 0.9 |
| 64 | 0.09108 | 0.1822 | 0.9089 |
| 65 | 0.08617 | 0.1723 | 0.9138 |
| 66 | 0.08331 | 0.1666 | 0.9167 |
| 67 | 0.0809 | 0.1618 | 0.9191 |
| 68 | 0.08568 | 0.1714 | 0.9143 |
| 69 | 0.09755 | 0.1951 | 0.9024 |
| 70 | 0.0896 | 0.1792 | 0.9104 |
| 71 | 0.07311 | 0.1462 | 0.9269 |
| 72 | 0.8955 | 0.2091 | 0.1045 |
| 73 | 0.9906 | 0.01877 | 0.009387 |
| 74 | 0.9959 | 0.008125 | 0.004062 |
| 75 | 0.9961 | 0.007759 | 0.00388 |
| 76 | 0.9961 | 0.007706 | 0.003853 |
| 77 | 0.9973 | 0.005361 | 0.00268 |
| 78 | 0.9999 | 0.0001718 | 8.592e-05 |
| 79 | 1 | 1.227e-05 | 6.134e-06 |
| 80 | 1 | 4.447e-06 | 2.224e-06 |
| 81 | 1 | 6.426e-06 | 3.213e-06 |
| 82 | 1 | 7.884e-06 | 3.942e-06 |
| 83 | 1 | 1.386e-05 | 6.932e-06 |
| 84 | 1 | 1.202e-05 | 6.01e-06 |
| 85 | 1 | 1.883e-05 | 9.413e-06 |
| 86 | 1 | 2.436e-05 | 1.218e-05 |
| 87 | 1 | 3.33e-05 | 1.665e-05 |
| 88 | 1 | 5.731e-05 | 2.866e-05 |
| 89 | 1 | 9.899e-05 | 4.949e-05 |
| 90 | 0.9999 | 0.00016 | 7.998e-05 |
| 91 | 1 | 3.224e-05 | 1.612e-05 |
| 92 | 1 | 4.747e-05 | 2.374e-05 |
| 93 | 1 | 7.281e-05 | 3.64e-05 |
| 94 | 0.9999 | 0.0001116 | 5.579e-05 |
| 95 | 1 | 9.878e-05 | 4.939e-05 |
| 96 | 1 | 6.693e-05 | 3.346e-05 |
| 97 | 1 | 1.345e-05 | 6.724e-06 |
| 98 | 1 | 1.583e-05 | 7.917e-06 |
| 99 | 1 | 2.882e-05 | 1.441e-05 |
| 100 | 1 | 4.691e-05 | 2.345e-05 |
| 101 | 1 | 6.617e-05 | 3.309e-05 |
| 102 | 0.9999 | 0.0001103 | 5.513e-05 |
| 103 | 0.9999 | 0.0001945 | 9.723e-05 |
| 104 | 0.9998 | 0.0003352 | 0.0001676 |
| 105 | 0.9997 | 0.0005701 | 0.000285 |
| 106 | 0.9996 | 0.0007449 | 0.0003724 |
| 107 | 0.9994 | 0.001168 | 0.0005839 |
| 108 | 0.9996 | 0.0008188 | 0.0004094 |
| 109 | 0.9994 | 0.001231 | 0.0006156 |
| 110 | 0.999 | 0.002069 | 0.001035 |
| 111 | 0.9983 | 0.003352 | 0.001676 |
| 112 | 0.9973 | 0.005403 | 0.002702 |
| 113 | 0.9962 | 0.007501 | 0.003751 |
| 114 | 0.9967 | 0.006576 | 0.003288 |
| 115 | 1 | 3.373e-06 | 1.686e-06 |
| 116 | 1 | 6.382e-06 | 3.191e-06 |
| 117 | 1 | 9.304e-06 | 4.652e-06 |
| 118 | 1 | 1.896e-05 | 9.478e-06 |
| 119 | 1 | 3.988e-05 | 1.994e-05 |
| 120 | 1 | 7.333e-05 | 3.666e-05 |
| 121 | 0.9999 | 0.0001336 | 6.681e-05 |
| 122 | 0.9999 | 0.0002565 | 0.0001282 |
| 123 | 0.9997 | 0.0005434 | 0.0002717 |
| 124 | 0.9996 | 0.0008452 | 0.0004226 |
| 125 | 0.9992 | 0.001699 | 0.0008495 |
| 126 | 0.9984 | 0.003274 | 0.001637 |
| 127 | 0.9974 | 0.005252 | 0.002626 |
| 128 | 0.9962 | 0.007565 | 0.003783 |
| 129 | 0.9963 | 0.007384 | 0.003692 |
| 130 | 0.9935 | 0.01293 | 0.006467 |
| 131 | 0.9884 | 0.02321 | 0.01161 |
| 132 | 0.9916 | 0.01675 | 0.008373 |
| 133 | 0.9844 | 0.03117 | 0.01558 |
| 134 | 0.9866 | 0.02679 | 0.01339 |
| 135 | 0.9778 | 0.04434 | 0.02217 |
| 136 | 0.9609 | 0.07828 | 0.03914 |
| 137 | 0.9251 | 0.1499 | 0.07494 |
| 138 | 0.8857 | 0.2286 | 0.1143 |
| 139 | 0.8317 | 0.3366 | 0.1683 |
| 140 | 0.7939 | 0.4122 | 0.2061 |
| 141 | 0.6468 | 0.7063 | 0.3532 |
| Meta Analysis of Goldfeld-Quandt test for Heteroskedasticity | |||
| Description | # significant tests | % significant tests | OK/NOK |
| 1% type I error level | 85 | 0.6855 | NOK |
| 5% type I error level | 101 | 0.814516 | NOK |
| 10% type I error level | 105 | 0.846774 | NOK |
| Ramsey RESET F-Test for powers (2 and 3) of fitted values |
> reset_test_fitted RESET test data: mylm RESET = 19.709, df1 = 2, df2 = 142, p-value = 2.794e-08 |
| Ramsey RESET F-Test for powers (2 and 3) of regressors |
> reset_test_regressors RESET test data: mylm RESET = 5.6964, df1 = 28, df2 = 116, p-value = 9.216e-12 |
| Ramsey RESET F-Test for powers (2 and 3) of principal components |
> reset_test_principal_components RESET test data: mylm RESET = 15.016, df1 = 2, df2 = 142, p-value = 1.214e-06 |
| Variance Inflation Factors (Multicollinearity) |
> vif
B C D E F G H I
1.559208 6.205108 8.256949 5.730235 2.693944 16.702143 9.086919 2.338467
J K L M N O
1.483640 2.522903 7.198786 1.953069 25.188350 22.830928
|