par4 <- '10'
par3 <- '0.1'
par2 <- '1'
par1 <- '0.1'
library(MASS)
PPCCBeta <- function(shape1, shape2, x)
{
x <- sort(x)
pp <- ppoints(x)
cor(qbeta(pp, shape1=shape1, shape2=shape2), x)
}
par1 <- as.numeric(par1)
par2 <- as.numeric(par2)
par3 <- as.numeric(par3)
par4 <- as.numeric(par4)
if (par1 < 0.1) par1 <- 0.1
if (par1 > 10) par1 <- 10
if (par2 < 0.1) par2 <- 0.1
if (par2 > 10) par2 <- 10
if (par3 < 0.1) par3 <- 0.1
if (par3 > 10) par3 <- 10
if (par4 < 0.1) par4 <- 0.1
if (par4 > 10) par4 <- 10
par1h <- par1*10
par2h <- par2*10
par3h <- par3*10
par4h <- par4*10
sortx <- sort(x)
c <- array(NA,dim=c(par2h,par4h))
for (i in par1h:par2h)
{
for (j in par3h:par4h)
{
c[i,j] <- cor(qbeta(ppoints(x), shape1=i/10,shape2=j/10),sortx)
}
}
bitmap(file='test1.png')
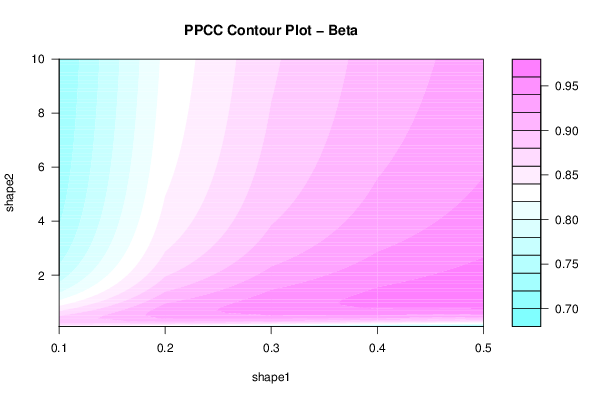
filled.contour((par1h:par2h)/10,(par3h:par4h)/10,c[par1h:par2h,par3h:par4h],xlab='shape1',ylab='shape2',main='PPCC Contour Plot - Beta')
dev.off()
xbar <- mean(x)
xvar <- var(x)
(a <- (xbar*(1-xbar)/xvar - 1)*xbar)
(b <- (1-xbar)*a/xbar)
(f<-fitdistr(x, 'beta',list(shape1=a,shape2=b)))
xlab <- paste('Beta(shape1=',round(f$estimate[[1]],2))
xlab <- paste(xlab,', shape2=')
xlab <- paste(xlab,round(f$estimate[[2]],2))
xlab <- paste(xlab,')')
bitmap(file='test2.png')
myser <- qbeta(ppoints(x), shape1=f$estimate[[1]], shape2=f$estimate[[2]])
qqplot(myser, x, main='QQ plot (Beta)', xlab=xlab )
grid()
dev.off()
load(file='createtable')
a<-table.start()
a<-table.row.start(a)
a<-table.element(a,'Parameter',1,TRUE)
a<-table.element(a,'Estimated Value',1,TRUE)
a<-table.element(a,'Standard Deviation',1,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'shape1',header=TRUE)
a<-table.element(a,f$estimate[1])
a<-table.element(a,f$sd[1])
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'shape2',header=TRUE)
a<-table.element(a,f$estimate[2])
a<-table.element(a,f$sd[2])
a<-table.row.end(a)
a<-table.end(a)
table.save(a,file='mytable.tab')
|